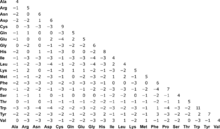
matrix substraction: MOLLI60 - BLOSUM62. Term by term substraction of... | Download Scientific Diagram

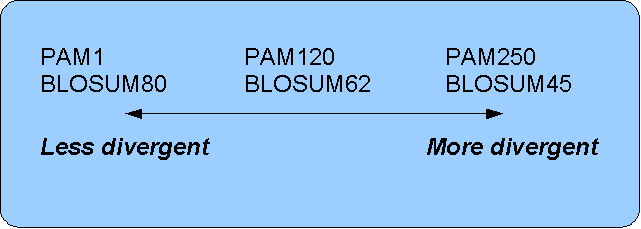
4.3 Scoring Matrices :: Chapter 4. Sequence Similarity :: Part II: Theory :: Basic local alignment search tool (blast) :: Misc :: eTutorials.org

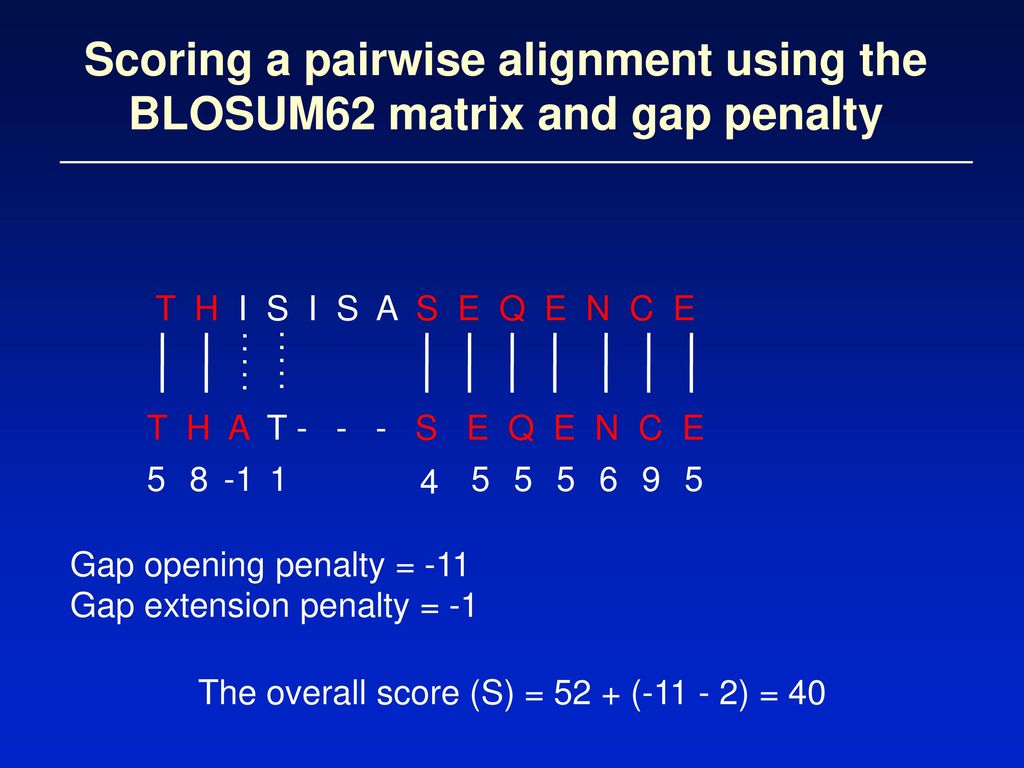
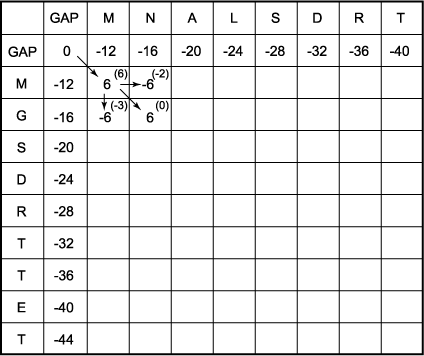
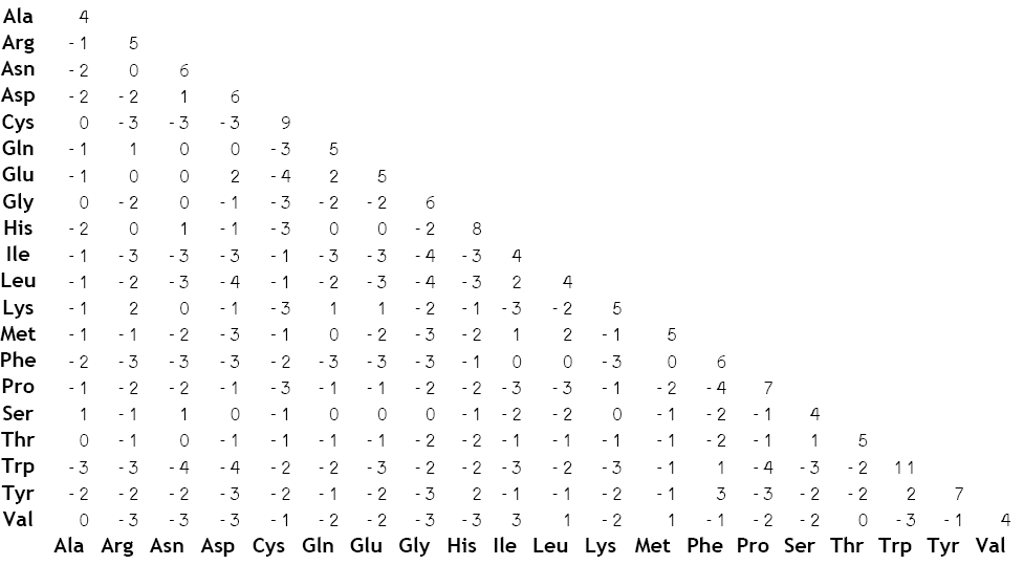
Learn about #BLOSUM-62 matrix | Calculate Max & Gap score for #BLAST-P with example using BLOSUM62 - YouTube

Comparison of 4 gap penalty schemes tested by pairwise BAliBASE scores... | Download Scientific Diagram
![PDF] Revisiting amino acid substitution matrices for identifying distantly related proteins | Semantic Scholar PDF] Revisiting amino acid substitution matrices for identifying distantly related proteins | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/b905b9bd1e1f1ed0bee8b8e986f6c3507928da10/3-Table1-1.png)
PDF] Revisiting amino acid substitution matrices for identifying distantly related proteins | Semantic Scholar

New amino acid substitution matrix brings sequence alignments into agreement with structure matches - Jia - 2021 - Proteins: Structure, Function, and Bioinformatics - Wiley Online Library

Learn about #BLOSUM-62 matrix | Calculate Max & Gap score for #BLAST-P with example using BLOSUM62 - YouTube